Here are 10 best practices that will improve how you structure your R projects, manage packages, organize files, and collaborate with others.
When installing packages, run install.packages() manually in the console- don’t include it in your scripts.
Installing a package is a one-time setup step, not part of your project’s workflow. Adding installation commands to a script only slows things down, since the check or installation would run every time the script is executed.
What should be included in your script is the library() call to load the package. This ensures the package is available whenever the script runs, and if it’s already loaded in the current session, R will simply skip it- so there’s no performance cost.
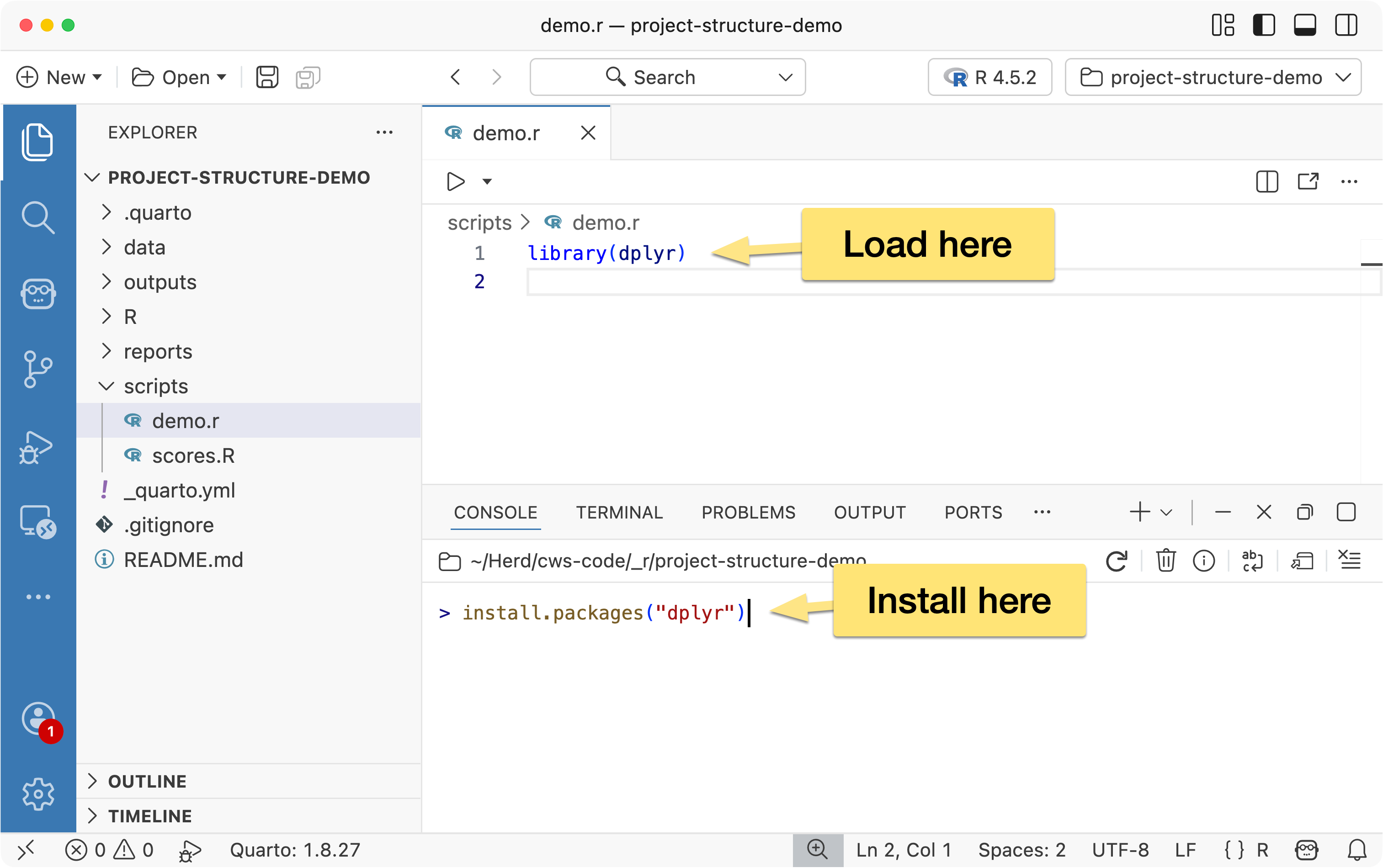
When loading packages, load only what you actually need. A common example is loading the entire tidyverse:
library(tidyverse)
…when the script only uses a couple of its packages:
library(dplyr)
library(ggplot2)
Being explicit about the packages you load keeps your environment cleaner, makes your dependencies clearer, and helps others understand exactly what your script relies on.
While we’re on the topic of packages, if you’re working on a complex project or collaborating with others, it’s worth using {renv} for package management.
{renv} installs packages on a project-by-project basis instead of relying on your system’s global library. This ensures each project uses the exact versions it needs, making your code more reproducible and allowing it to run consistently across different machines.
One of the simplest workflow improvements you can make in R is adopting a consistent project structure.
Rather than scattering files across your computer organize each project in a clear, logical way, separating components like data, scripts, reports, and output.
This makes your work easier to understand, reproduce, and share.
Here’s a suggested outline:
project/
data/
R/
scripts/
outputs/
reports/
README.md
→ Learn more about project structure best practices...
In most programming workflows, it’s standard practice to include a README.md file at the root of your project. This file acts as a high-level introduction and guide for anyone working with the project — including your future self.
A good README.md typically includes:
The goal of the README is to help someone quickly understand what the project does, how it’s organized, and how to get started — without having to dig through the source code.
As R projects grow, it becomes harder to manage all the steps involved in your analysis — importing data, cleaning it, modeling, visualizing, and generating reports. The {targets} package helps you organize these steps into a clear, reproducible workflow.
Instead of running scripts manually from top to bottom, {targets} lets you define your project as a series of connected steps, where each step depends on the results of previous steps.
→ Learn more about {targets}...
When sourcing files, avoid using absolute, machine-specific paths, as these will break when someone else runs your code on their machine.
Instead, use relative paths that locate files within the project directory. This keeps your projects portable, easier to share, and more reproducible.

When working with sensitive information such as API keys, database credentials, or access tokens, you should avoid placing these values directly in your scripts. Instead, store them in a .Renviron file and access them within your code using Sys.getenv(). Unlike your script files, .Renviron is excluded from version control.

This approach ensures that sensitive credentials are never committed to your repository and allows each collaborator to manage their own environment variables securely on their local machine.
→ Learn more about R environment variables...
Understanding how to use version control is an essential skill for any programmer, including those working in R and data science.
Version control allows you to track changes, back up your work, and collaborate more safely. Even on solo projects, it gives you the freedom to experiment without worrying about breaking things.
You don’t need to be a Git expert — even basic use offers significant benefits in traceability, organization, and long-term project management.
→ Learn about Git Version Control in RStudio...
→ Learn about Git Version Control in Positron...
Raw data should be treated as read-only.
Once you collect, download, or receive your original data files, do not modify them directly. Instead, keep them unchanged in a dedicated data/ folder and perform all cleaning, transformation, and analysis steps in scripts.